The Whole C9 Yards Explore our 2014 timeline to learn more about emerging strategies to diagnose and treat C9 ALS.
Since the discovery of C9orf72 in 2012, the most common form of sporadic and familial ALS identified to date, potential underlying mechanisms are unraveling at a rapid pace. And, potential strategies to diagnose and treat this form of the disease are beginning to emerge.
In just the last two years alone, a genetic test for C9orf72 ALS (C9 ALS) entered the clinic.
A PET imaging technique, developed by Katholieke Universiteit Leuven’s Koen Van Laere MD PhD and Università di Torino’s Adriano Chiò MD, emerged – opening the door to predicting the outcome of people with C9 ALS. An MRI method, developed by Trinity College Dublin’s Peter Bede MD, which aims to help anticipate cognitive challenges, surfaced. And, a new map charting the spread of C9 ALS created by the research teams of Universitätsklinikum Ulm’s Heiko Braak MD and University of Pennsylvania’s John Trojanowski MD PhD may help clinicians in the future predict the course of their patients' disease.
In addition, the first genetic modifier of C9 ALS, TMEM 106B, was discovered – paving the way toward a genetic test for C9 ALS-FTD.
“This is one of the first modifiers that could really explain why some people get ALS and FTD,” says Mayo Clinic’s Rosa Rademakers PhD.
All about RNA?
A potential antisense therapy, which aims to reduce levels of possibly toxic repeat-rich C9 RNAs, is in the preclinical stage. And, an initiative to discover RNA-targeted small molecules led by Scripps Institute’s Matthew Disney PhD launched.
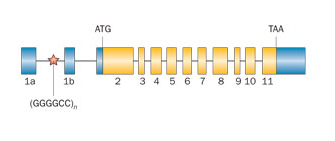
Don't repeat that Nearly 40% of inherited cases of ALS appear to be caused, at least in part, by expanded repeat sequences in the C9orf72 gene. Image: Courtesy of Nature Publishing Group.
Key questions remain. Expanded C9 RNAs appear to make motor neurons more vulnerable to ALS according to studies led by Jeff Rothstein MD PhD. But whether the reduction of these RNAs alone is sufficient to treat people with C9 ALS remains an open question.
Toxic proteins also may contribute to the destruction of motor neurons according to studies led by Ludwig Maximilians Universität München’s Christian Haass PhD and Mayo Clinic’s Len Petrucelli PhD. And, according to studies led by La Trobe University’s Julie Atkin PhD, the reduction of C9orf72 protein, may in part make the removal of misfolded proteins more difficult – further contributing to the disease.
What’s more, existing mouse models of the disease do not appear to exhibit key symptoms of ALS according to University of Massachusetts’ Bob Brown MD PhD– making the development of potential therapies challenging.
"It is important to keep striving for models that actually recaptiulate the pathology, because then you start to learn which pieces of the C9 puzzle actually lead to the disease," says ALS Therapy Development Institute's Fernando Vieira MD.
Testing the limits
The utility of genetic tests also remains limited.
People at high risk of developing C9 ALS can be detected by existing methods. But identifying people who will develop the disease remains tricky to do.
The reason, at least in part, is that C9 ALS appears to be one of a growing number of forms of ALS that may be oligogenic in nature. Multiple mutations in multiple genes may contribute to the onset and progression of the disease.
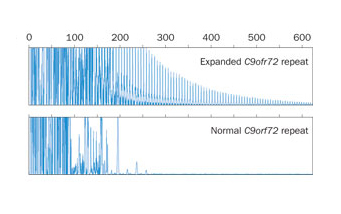
Standardized tests? Existing blood tests help identify people with C9 ALS by detecting expanded repeat-rich RNAs. But according to a study led by University of Umeå's Peter Andersen MD, nearly 8% of people may diagnosed incorrectly due to a lack of testing standards. Image: Courtesy of Nature Publishing Group.
An alteration in at least one other gene linked to ALS can be detected in people with C9 ALS - at least in some cases according to results from Marka Blitterswijk MD PhD, now at Mayo Clinic in Florida.
“It is probably the tip of the iceberg,” says University of Massachusetts’ Bob Brown MD PhD. “It is clear that we can find more than one variant in a person with ALS.”
What’s more, existing genetic tests only help identify people with C9 ALS. The size of the expanded RNA – at least in key cells isolated from the blood and the skin – do not appear to correlate with the progression rate of their disease.
“There is little we can do with repeat length to predict clinical outcome,” says Rosa Rademakers PhD.
***
To learn more about C9orf72 ALS, check out ALS Antisense and Sensibility. To find out more about emerging diagnostic strategies for the disease, check out C9 Comes Into Focus.
References
Akimoto, C. et al. (2014) A blinded international study on the reliability of genetic testing for GGGGCC-repeat expansions in C9orf72 reveals marked differences in results among 14 laboratories. Journal of Medical Genetics doi:10.1136/jmedgenet-2014-102360. Abstract | Full Text
Haeusler, A.R. et al. (2014) C9orf72 nucleotide repeat structures initiate molecular cascades of disease. Nature 507(7491), 195-200. Abstract | Full Text (Subscription Required)
van Blitterswijk, M. et al. (2014) TMEM106B protects C9ORF72 expansion carriers against frontotemporal dementia. Acta Neuropathologica 127(3), 397-406. Abstract | Full Text (Subscription Required)
Gallagher, M.D. et al. (2014) TMEM106B is a genetic modifier of frontotemporal lobar degeneration with C9orf72 hexanucleotide repeat expansions. Acta Neuropathologica 127(3), 407-418. Abstract | Full Text (Subscription Required)
Cistaro, A. et al. (2014) The metabolic signature of C9ORF72-related ALS: FDG PET comparison with nonmutated patients. European Journal of Nuclear and Molecular Imaging doi: 10.1007/s00259-013-2667-5 Abstract | Full Text (Subscription Required)
Van Laere, K., Vanhee, A., Verschueren, J., De Coster, L., Driesen, A., Dupont, P., Robberecht, W. and Van Damme, P. (2014) Value of 18Fluorodeoxyglucose-Positron-Emission Tomography in Amyotrophic Lateral Sclerosis: A Prospective Study. JAMA Neurology doi: 10.1001/jamaneurol.2014.62. Abstract | Full Text (Subscription Required)
Farg, M.A. et al. (2014) C9ORF72, implicated in amytrophic lateral sclerosis and frontotemporal dementia, regulates endosomal trafficking. Human Molecular Genetics doi: 10.1093/hmg/ddu068 Abstract | Full Text
Waite, A.J., Bäumer, D., East, S., Neal, J., Morris, H.R., Ansorge, O., Blake and D.J. (2014) Reduced C9orf72 protein levels in frontal cortex of amyotrophic lateral sclerosis and frontotemporal degeneration brain with the C9ORF72 hexanucleotide repeat expansion. Neurobiology of Aging doi: 10.1016/j.neurobiolaging.2014.01.016. Abstract | Full Text (Subscription Required)
Lee, Y.B. et al. (2013) Hexanucleotide repeats in ALS/FTD form length-dependent RNA foci, sequester RNA binding proteins, and are neurotoxic. Cell Reports 5(5), 1178-1186.Abstract | Full Text
Donnelly, C.J. et al. (2013) RNA toxicity from the ALS/FTD C9ORF72 expansion is mitigated by antisense intervention. Neuron 80(2), 415-428. Abstract | Full Text (Subscription Required)
Sareen, D. et al. (2013) Targeting RNA foci in iPSC-derived motor neurons from ALS patients with a C9ORF72 repeat expansion. Science Translational Medicine 5(208), 208ra149. Abstract | Full Text (Subscription Required)
Lagier-Tourenne, C. et al. (2013) Targeted degradation of sense and antisense C9orf72 RNA foci as therapy for ALS and frontotemporal degeneration. Proceedings of the National Academy of Sciences 110(47), E4530-E4539. Abstract | Full Text (Subscription Required)
Almeida, S. et al. (2013) Modeling key pathological features of frontotemporal dementia with C9ORF72 repeat expansion in iPSC-derived human neurons. Acta Neuropathologica 126(3), 385-399. Abstract | Full Text
van Blitterswijk, M. et al. (2013) C9ORF72 repeat expansions in cases with previously identified pathogenic mutations. Neurology 81(15), 1332-1341. Abstract | Full Text
van Blitterswijk, M. et al. (2013) Association between repeat sizes and clinical and pathological characteristics in carriers of C9ORF72 repeat expansions (Xpansize-72): a cross-sectional cohort study. Lancet Neurology 12(10), 978-988.Abstract | Full Text (Subscription Required)
Panda, S.K., Wefers, B., Ortiz, O., Floss, T., Schmid, B., Haass, C., Wurst, W. and Kühn, R. (2013) Highly efficient targeted mutagenesis in mice using TALENs. Genetics 195(3), 703-713. Abstract | Full Text (Subscription Required)
Brettschneider, J. et al. (2013) Stages of pTDP-43 pathology in amyotrophic lateral sclerosis. Annals of Neurology 74(1), 20-38. Abstract | Full Text (Subscription Required)
Bede, P., Bokde, A.L., Byrne, S., Elamin, M., McLaughlin, R.L., Kenna, K., Fagan, A.J., Pender, N., Bradley, D.G. and Hardiman, O. (2013) Multiparametric MRI study of ALS stratified for the C9orf72 genotype. Neurology 81(4), 361-369. Abstract | Full Text (Subscription Required)
Xi, Z. et al. (2013) Hypermethylation of the CpG island near the G4C2 repeat in ALS with a C9orf72 expansion. American Journal of Human Genetics 92(6), 981-989. Abstract | Full Text
Xu, Z. et al. (2013) Expanded GGGGCC repeat RNA associated with amyotrophic lateral sclerosis and frontotemporal dementia causes neurodegeneration. Proceedings of the National Academy of Sciences 110(19), 7778-7783. Abstract | Full Text
Mori, K. et al. (2013) hnRNP A3 binds to GGGGCC repeats and is a constituent of p62-positive/TDP43-negative inclusions in the hippocampus of patients with C9orf72 mutations.Acta Neuropathologica 125(3), 1178-1186. Abstract |Full Text (Subscription Required)
Mori, K. et al. (2013) The C9orf72 GGGGCC repeat is translated into aggregating dipeptide-repeat proteins in FTLD/ALS. Science 339(6125), 1335-1338. Abstract | Full Text (Subscription Required)
Ash, P.E. et al. (2013) Unconventional translation of C9ORF72 GGGGCC expansion generates insoluble polypeptides specific to c9FTD/ALS. Neuron 77(4) 639-646. Abstract | Full Text (Subscription Required)
Williams, K.L., Fifita, J.A., Vucic, S., Durnall, J.C., Kiernan, M.C., Blair, I.P. and Nicholson, G.A. (2013) Pathophysiological insights into ALS with C9ORF72 expansions. Journal of Neurology, Neurosurgery and Psychiatry 84(8), 931-935. Abstract | Full Text (Subscription Required)
Gómez-Tortosa, E. et al. (2013) C9ORF72 hexanucleotide expansions of 20-22 repeats are associated with frontotemporal deterioration. Neurology 80(4), 366-370. Abstract | Full Text (Subscription Required)
Reddy, K., Zamiri, B., Stanley, S.Y., Macgregor, R.B. Jr and Pearson, C.E. (2013) The disease-associated r(GGGGCC)n repeat from the C9orf72 gene forms tract length-dependent uni- and multimolecular RNA G-quadruplex structures. Journal of Biological Chemistry 288(14), 9860-9866. Abstract | Full Text
Fratta, P., Mizielinska, S., Nicoll, A.J., Zloh, M., Fisher, E.M., Parkinson, G. and Isaacs, A.M. (2012) C9orf72 hexanucleotide repeat associated with amyotrophic lateral sclerosis and frontotemporal dementia forms RNA G-quadruplexes. Scientific Reports 2, 2016. Abstract | Full Text
van Blitterswijk, M. et al. (2012) Evidence for an oligogenic basis of amyotrophic lateral sclerosis. Human Molecular Genetics 21(17), 3776-3784. Abstract | Full Text
Majounie, E., et al. (2012) Frequency of the C9orf72 hexanucleotide repeat expansion in patients with amyotrophic lateral sclerosis and frontotemporal dementia: a cross-sectional study. Lancet Neurology 11(4), 323-330. Abstract | Full Text
DeJesus-Hernandez, M., et al. (2011) Expanded GGGGCC hexanucleotide repeat in noncoding region of C9ORF72 causes chromosome 9p-linked FTD and ALS. Neuron 72(2), 245-256. Abstract | Full Text
Renton, A. E., et al. (2011) A hexanucleotide repeat expansion in C9ORF72 is the cause of chromosome 9p21-linked ALS-FTD. Neuron 72(2), 257-268. Abstract | Full Text
Vance, C. et al. (2006) Familial amyotrophic lateral sclerosis with frontotemporal dementia is linked to a locus on chromosome 9p13.2-21.3. Brain 129(4), 868-876. Abstract | Full Text