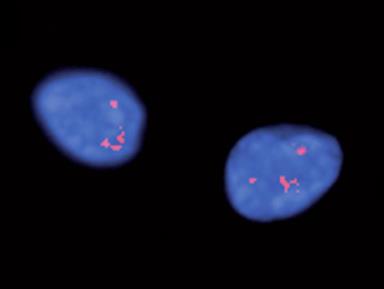
Repeat offenders Expanded RNAs appear to tightly fold up (RNA foci, above) in the nuclei of motor neurons in people with C9orf72-linked ALS - keeping key regulatory proteins out of action. Image: Courtesy of Nature Publishing Group.
With the discovery of disruptions in C9orf72 in 2011, ALS began to emerge as a whole brain disease. Key network glitches, detected by MRI, could be traced back to structural changes in cognitive centers of the brain – potentially explaining key cognitive challenges including problem solving and complex decision making (executive function). And, key behavioral signs including apathy and assessing risk, characteristic of frontotemporal dementia (FTD), became more widely appreciated in people with ALS – further blurring the line between these diseases.
“We used to think of ALS and FTD as different diseases. But now it is becoming clear that it is a disease continuum,” says University of Leuven’s Wim Robberecht MD PhD.
C9orf72 is now one of ten genes implicated in both ALS and ALS/FTD according to University of Massachusetts’ Bob Brown MD PhD. And, repeat expansions in the gene appear to be linked to at least 1 out of 20 cases of ALS – including more than 40% of people with familial ALS (fALS) – across the globe.
“This is clearly the most common cause of neurodegeneration,” says Brown.
Key questions remain. Expanded RNAs appear to accumulate in people with C9orf72-linked ALS – potentially keeping regulatory proteins out on the cytoplasmic sidelines. But according to recent studies led by Deutsches Zentrum für Neurodegenerative Erkrankungen’s Christian Haass PhD and Mayo Clinic’s Len Petrucelli PhD, the disease may instead be a perfect storm of proteins and RNAs brewing in people with the disease. Repeat-associated proteins called C9RANTs also appear to build up in motor neurons in people with C9orf72-linked ALS – potentially fueling their destruction. And, the production of C9orf72 protein appears to be reduced in people with C9orf72-linked ALS and may also contribute to the disease.
But research teams led by University of California San Diego’s Don Cleveland PhD and Johns Hopkins University School of Medicine's Jeff Rothstein MD PhD remain undaunted – developing treatment strategies that aim to destroy expanded RNAs in people with the disease.
The Whole C9 Yards Explore our interactive timeline to learn more about C9orf72, the most common form of ALS identified to date.
Ahead of the 2013 meeting of the Society of Neuroscience, researchers gathered at the RNA Metabolism and Neurological Disease conference in San Diego to talk about the latest advances in C9orf72-linked ALS and emerging treatment strategies for the disease going forward.
Modeling workshop
The race is on to create a model of C9orf72-linked ALS – a key first step in generating a treatment strategy for the disease.
University of Massachusetts’ Bob Brown MD PhD is leading the pack - developing a mouse model of the disease.
The transgenic mice, which harbors 580 repeats in the first intron of the C9orf72 gene, do not appear to develop paralysis. But according to Brown, may have other emerging hallmarks of the disease.
“It is early days,” says Brown.
But whether these mice actually exhibit key symptoms of ALS makes no difference according to University of California San Diego’s Don Cleveland PhD, who is also developing them.
“We don’t care whether these mice develop full or partial disease,” says Cleveland. “It is not a key feature to move forward.”
Researchers, however, need to keep trying to develop mouse models of C9orf72-linked ALS, says ALS Therapy Development Institute’s Fernando Vieira MD.
Although these mice will be helpful in validating emerging antisense treatment strategies for C9orf72-linked ALS, says Vieira, we don’t know what drives the disease. And, therefore we cannot be sure which therapy to develop.
“It is important to keep striving for models that actually recapitulate elements of ALS symptomatology,” says Vieira. “Then, we start to learn which pieces of the C9orf72 puzzle actually lead to the disease.”
Meanwhile, Johns Hopkins University School of Medicine’s Jeff Rothstein MD PhD is turning to stem cells to develop a treatment strategy for C9Orf72-linked ALS.
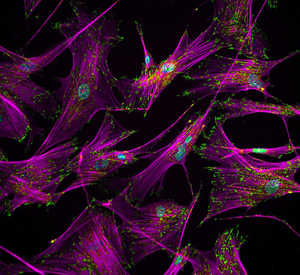
Skin to skin A growing number of scientists are turning to stem cells, recreated from skin cells (above) from people with C9orf72-linked ALS, in hopes to discover more effective medicines for the disease. Image: Ankur Singh PhD, Georgia Institute of Technology.
Motor neurons are recreated from people with ALS by reprogramming their skin cells. And, the cells scoured for key signs of disease.
The approach aims to identify key phenotypes that can be used to develop medicines that help weather these motor neurons from the C9orf72 storm by detoxifying them.
The strategy is the same strategybeing developed by Cedar Sinai’s Bob Baloh MD PhD and University of California San Diego’s Clotilde Lagier-Tourenne PhD.
“We do not need a mouse to move a treatment forward,” says Rothstein.
Motor neurons do appear to exhibit tell tale signs of disease. Expanded C9orf72 RNAs and C9RANTs can be detected according to Mayo Clinic's Len Petrucelli PhD and University of Florida College of Medicine's Laura Ranum PhD. And, according to Johns Hopkins University School of Medicine’s Chris Donnelly PhD, the motor neurons may also be more vulnerable – at least to increased levels of glutamate which occurs in many people with the disease.
But whether this model alone can be used to develop treatments for the disease remains an open question according to Mayo Clinic’s Rosa Rademakers PhD. Thousands of repeats appear to be embedded in the C9orf72 gene in key cells in the brain and spinal cord. But only at most 700 – 800 repeats can be detected in skin cells – the cells researchers use to model C9orf72-linked disease.
“I think it is extremely important to keep this in mind,” says Rademakers.
Storm Troopers
In 2011, research teams led by Mayo Clinic’s Rosa Rademakers PhD and National Institutes of Health’s Bryan Traynor MD PhD fingered toxic RNA as the prime suspect behind C9orf72-linked ALS.
Repeat-rich RNAs appear to be detected in motor neurons of people with C9orf72-linked ALS. And, suspected researchers, kept key regulatory proteins out of commission – leading to the disease.
The same underlying mechanism that leads to progressive muscle weakness in people with myotonic dystrophy.
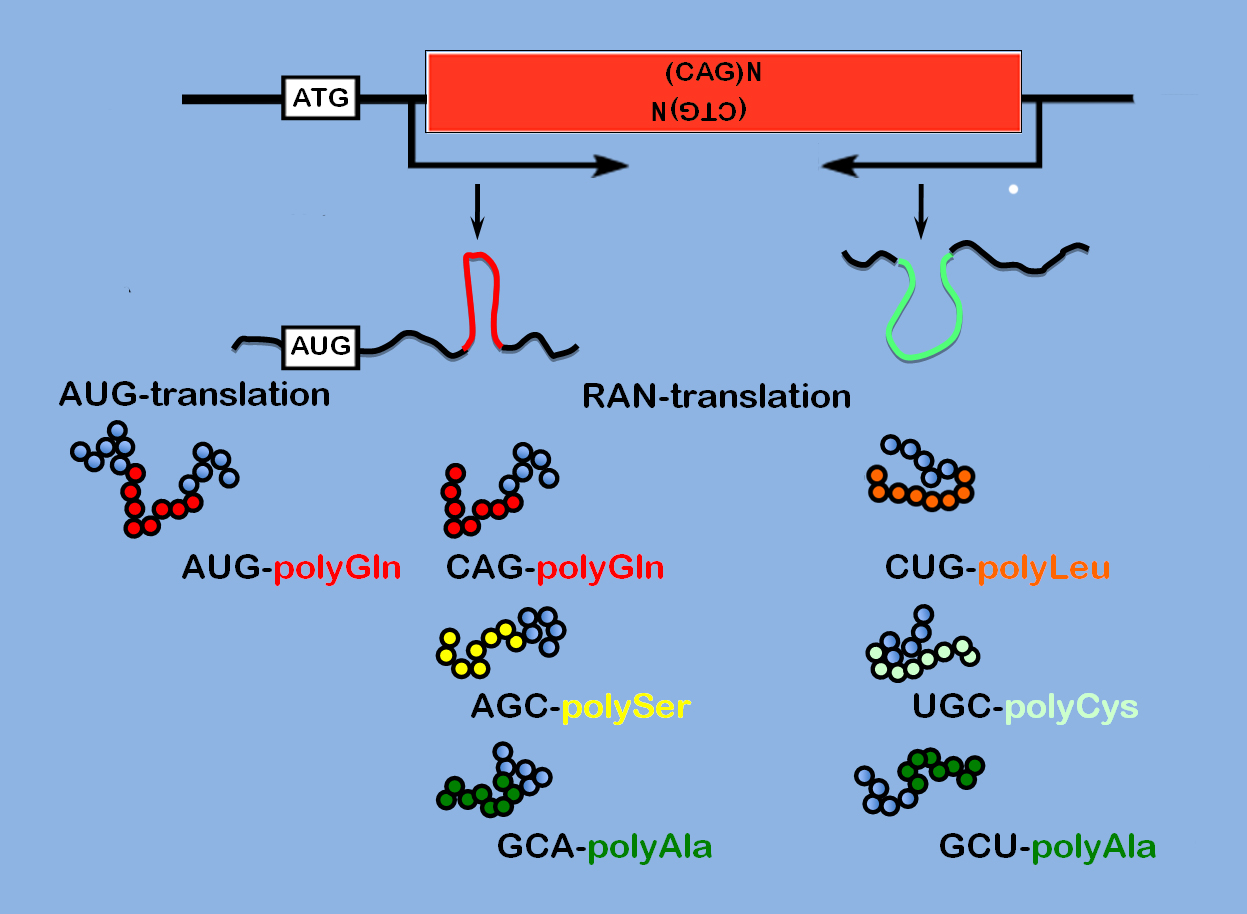
RANT and rave? RNAs appear to tightly fold up in people with C9orf72-linked ALS - enabling the synthesis of potentially toxic short proteins called C9orf72 non-ATG translated peptides (C9RANTs). RANTs are emerging as a potentially key feature in a growing number of diseases including myotonic dystrophy (above). Image: Courtesy of the Public Library of Science.
With the discovery of short dipeptide repeat proteins in people with C9orf72-linked ALS in 2012, however, researchers began to realize that the disease is much more complicated.
As many as six dipeptides appear to accumulate in key neurons in the brain in people with C9orf72-linked ALS and ALS/FTD according to studies led by University of Florida College of Medicine’s Laura Ranum PhD. And, at least one of these short proteins according to cell culture studies presented by Mayo Clinic’s Len Petrucelli PhD may damage them.
The short dipeptides are emerging as a characteristic feature of a growing number of RNA repeat-associated diseases including myotonic dystrophy I (DM1) and fragile X-associated tremor/ataxia syndrome (FXTAS).
“We need to start thinking about RAN translation,” says University of Florida College of Medicine’s Laura Ranum PhD. “And, their role in these diseases.”
But although there are a lot of lessons that can be learned from these RNA repeat-associated diseases, says Hospital of Sick Kids’ Christopher Pearson PhD, researchers still need to prove that the same underlying mechanisms lead to ALS.
At least 7 regulatory proteins appear to be kept out of action by C9orf72 expanded RNAs in the brain and spinal cord according to Johns Hopkins University School of Medicine’s Chris Donnelly PhD. But whether the loss of any of them contributes to ALS remains an open question.
What’s more, researchers remain unsure what role if any C9RANTs play in the disease.
“Toxicity is a question that is looming over all of us,” says Johns Hopkins School of Medicine’s Jeff Rothstein MD PhD.
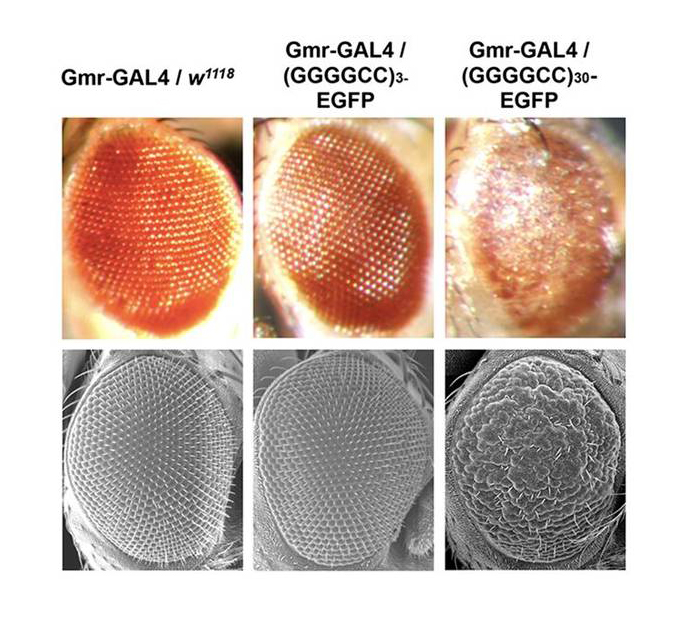
Eye of the C9 Storm Researchers hope to identify key proteins that rescue the rough eye phenotype (top, right) of C9orf72 fruit flies in hopes to unravel underlying mechanisms of C9orf72-linked ALS. Image: Courtesy of the National Academy of Sciences.
To begin to tackle this question, the Johns Hopkins University School of Medicine team is turning to an emerging model of C9orf72-linked ALS in hopes to identify key sequestered proteins that may contribute to the disease.
The transgenic fruit flies, which produce expanded repeats in key regions of the eye, sustain a progressive loss of key ocular neurons which leads to a “rough” appearance.
The team hopes to identify key proteins benched by repeat-rich C9orf72 RNAs by increasing their levels in the eye. And, looking to see whether their eyes are restored.
“Some RNA-binding proteins appear to reduce the rough eye phenotype,” says Johns Hopkins’ Christopher Donnelly PhD.
The GAL4 UAS fruit fly model, developed by Zihui Xu PhD, now at Huazhong University of Science and Technology, is one of the first to recreate neuron loss, a key feature of the disease.
Meanwhile, scientists including Mayo Clinic’s Len Petrucelli PhD are trying to figure out whether dipeptide repeat proteins also contribute to the disease.
“We are trying to understand what C9RANTs are doing,” says Mayo Clinic’s Len Petrucelli PhD, “whether they are toxic.”
Target Practice
A potential treatment strategy for C9orf72-linked ALS appears to be safe and well tolerated – at least in mice according to University of California San Diego’s Clotilde Lagier-Tourenne PhD.
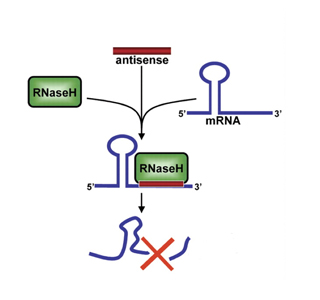
Search and destroy Antisense oligonucleotides directed against the repeat expanded form of C9orf72 RNA bind these potentially toxic molecules- tagging them for destruction. Image: Courtesy of Rockefeller University Press.
The approach, developed by ISIS Pharmaceuticals, deploys antisense oligonucleotides (ASOs) known as gapmers to search for expanded C9orf72 RNAs and target them for destruction.
The emerging treatment strategy does appear to reduce levels of repeat-rich C9orf72 RNAs in motor neurons recreated from people with the disease.
Treatment with antisense oligonucleotides appear to reduce levels of expanded C9orf72 RNAs (RNA foci) to nearly 50% according to University of California San Diego’s Clotilde Lagier-Tourenne PhD. And, according to Johns Hopkins University School of Medicine’s Chris Donnelly PhD, may help protect them from potentially toxic buildup of glutamate (excitotoxicity) – a potential contributor to the disease.
What's more, many of these antisense oligonucleotides do not appear to further reduce expression of the C9orf72 gene.
The results add to growing evidence that this emerging treatment strategy may be helpful to people with C9orf72-linked disease.
“It is really encouraging – going forward with this strategy in C9orf72 ALS,” says Lagier-Tourenne.
But whether antisense is the best approach to treat C9orf72-linked ALS remains an open question. Existing dipeptide repeat proteins remain untouched. Expanded RNAs tightly fold up into stable quadruplexes according to studies led by Hospital of Sick Kids’ Christopher Pearson PhD – making them potentially less vulnerable to oligo attack. And, the battle may need to be waged in the nervous system. ASOs are deployed intrathecally – directly into the spinal cord.
Small molecules may be a better choice to keep key regulatory proteins in action according to Scripps Institute’s Matthew Disney PhD. Small molecules can often be taken by mouth. And, may bind more tightly – burying themselves in the expanded RNA’s unique folds.
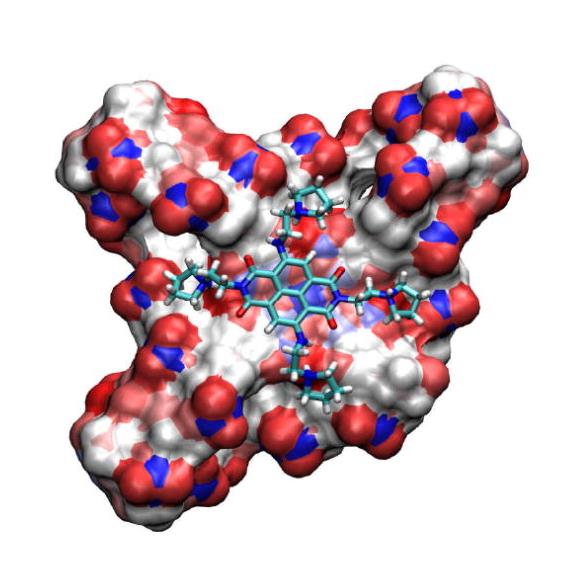
Foursquare Researchers hope to design small molecules that hide out in the unique folds of the expanded C9orf72 RNA 'quadruplexes' in people with ALS -to keep regulatory proteins in action. Image: Reprinted with permission from Gunaratnam, M. et al. (2009). Copyright 2009 American Chemical Society.
“We are targeting secondary structure,” explains Disney. “This is more selective than oligonucleotides.”
But designing small molecules that target these RNAs may be tricky to do. Small molecule libraries contain few compounds that bind RNAs according to Disney. And, no antimicrobial or antiviral substance (natural product) has been discovered to date that target RNAs - providing no clues to create them.
To overcome these challenges, the Scripps Institute team introduced a new method to design molecules that target expanded RNAs to treat disease.
The library vs library approach involves the identification of key small molecules that target the RNA. And, tweaking them using computational-based methods to optimize their abilities to bind and detoxify them.
The resulting small molecules can then be displayed on peptide-like scaffolds to enable them to bind cooperatively to the repeat-rich RNAs – maximizing their efficacy.
The approach is currently being used to develop potential treatments for rare muscles diseases known as myotonic dystrophies.
Efforts to create expanded C9orf72 repeat-rich RNA-targeted small molecules are now underway.
***
To learn more about antisense and its potential for people with C9orf72-linked ALS, tune into our podcast with Jeff Rothstein MD PhD. To find out more about C9orf72-linked ALS and potential stem cell treatments for the disease, check out Astrocytes for the people?
References
Donnelly, C.J. et al. (2013) RNA toxicity from the ALS/FTD C9ORF72 expansion is mitigated by antisense intervention. Neuron 80(2), 415-420. Abstract | Full Text (Subscription Required)
Lagier-Tourenne, C. et al. (2013) Targeted degradation of sense and antisense C9orf72 RNA foci as therapy for ALS and frontotemporal degeneration. Proceedings of the National Academy of Sciences 110(47), E4530-4539. Abstract | Full text (Subscription Required)
Sareen, D. et al. (2013) Targeting RNA foci in iPSC-derived motor neurons from ALS patients with a C9ORF72 repeat expansion. Science Translational Medicine 5(208), 208ra149. Abstract | Full Text (Subscription Required)
Zu, T. et al. (2013) RAN proteins and RNA foci from antisense transcripts in C9ORF72 ALS and frontotemporal dementia. Proceedings of the National Academy of Sciences 110(51), E4968-E4977. Abstract | Full Text
Mori, K. et al. (2013) The C9orf72 GGGGCC repeat is translated into aggregating dipeptide-repeat proteins in FTLD/ALS. Science 339(6125), 1335 – 1338. Abstract | Full Text (Subscription Required)
Ash, P.E. et al. (2013) Unconventional translation of C9ORF72 GGGGCC expansion generates insoluble polypeptides specific to c9FTD/ALS. Neuron 77(4), 639-646. Abstract | Full Text (Subscription Required)
Xu, Z. et al. (2013) Expanded GGGGCC repeat RNA associated with amyotrophic lateral sclerosis and frontotemporal dementia causes neurodegeneration. Proceedings of the National Academy of Sciences 110(19), 7778–7783. Abstract | Full Text
Reddy, K., Zamiri, B., Stanley, S.Y., Macgregor, R.B. Jr and Pearson, C.E. (2013) The disease-associated r(GGGGCC)n repeat from the C9orf72 gene forms tract length-dependent uni- and multimolecular RNA G-quadruplex structures. Journal of Biological Chemistry 288(14), 9860-9866.Abstract|Full Text (Subscription Required)
Majounie, E. et al. (2012) Frequency of the C9orf72 hexanucleotide repeat expansion in patients with amyotrophic lateral sclerosis and frontotemporal dementia: a cross-sectional study. Lancet Neurology 11(4):323-30. Abstract | Full Text
DeJesus-Hernandez, M. et al. (2011) Expanded GGGGCC hexanucleotide repeat in a noncoding region of C9ORF72 causes chromosome 9p-linked FTD and ALS Neuron 72(2), 245-256. Abstract | Full Text (Subscription Required)
Renton, A.E. et al. (2011) A hexanucleotide repeat expansion in C9ORF72 is the cause of chromosome 9p21-linked ALS-FTD. Neuron 72(2), 257-268. Abstract | Full Text (Subscription Required)
Further Reading
van Blitterswijk, M., DeJesus-Hernandez, M. and Rademakers R. (2012) How do C9ORF72 repeat expansions cause amyotrophic lateral sclerosis and frontotemporal dementia: can we learn from other noncoding repeat expansion disorders? Current Opinion in Neurology 25(6), 689-700. Abstract | Full Text (Subscription Required)
Disney, M.D. (2013) Rational design of chemical genetic probes of RNA function and lead therapeutics targeting repeating transcripts. Drug Discovery Today 18(23-24), 1228-1236. Abstract | Full Text (Subscription Required)